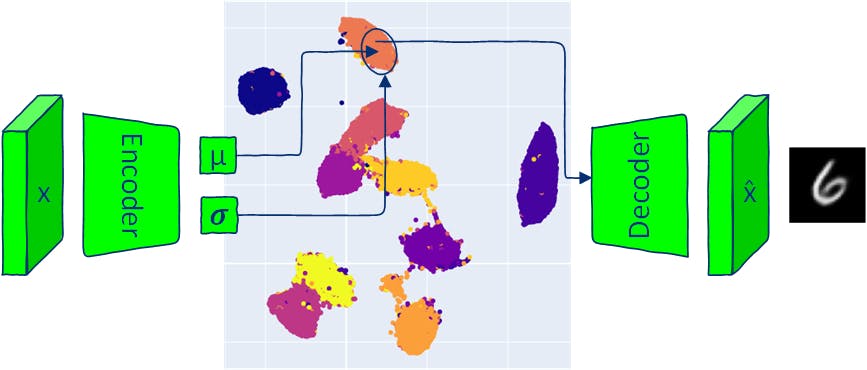
Same as traditional autoencoders, VAE architecture has two parts: an encoder and a decoder. Traditional AE models map inputs into a latent-space vector and reconstruct the output from this vector.
VAE maps inputs into a multivariate normal distribution (the encoder outputs the mean and the variance of each latent dimension).
Since the VAE encoder produces a distribution, the new data can be generated by sampling from this distribution and passing the sampled latent vector into the decoder. Sampling from produced distribution to generate output images means that VAE allows the generating of novel data that is similar, but identical to the input data.
This article explores components of VAE architecture and provides several ways of generating new images (sampling) with VAE models. All the code is available in Google Colab.
1 VAE Model Implementation
AE model is trained by minimizing reconstruction loss (for example BCE or MSE)
Autoencoders and Variational Autoencoders both have two parts: encoder and decoder. The encoder neural network of AE learns to map each image into a single vector in latent space and the decoder learns to reconstruct the original image from the encoded latent vector.
VAE model is trained by minimizing reconstruction loss and KL-divergence loss
The encoder neural network of VAE outputs parameters that define a probability distribution for each dimension of the latent space (multivariate distribution). For each input, the encoder produces a mean and a variance for each dimension of latent space.
The output mean and variance are used to define a multivariate Gaussian distribution. The decoder neural network is the same as in AE models.
1.1 VAE Losses
The goal of training a VAE model is to maximize the likelihood of generating real images from provided latent vectors. During training, the VAE model minimizes two losses:
- reconstruction loss - the difference between the input images and the output of the decoder.
- Kullback–Leibler divergence loss (KL Divergence a statistic distance between two probability distributions) - the distance between the probability distribution of the encoder's output and a prior distribution (a standard normal distribution), helping to regularize the latent space.
1.2 Reconstruction Loss
Common reconstruction losses are binary cross-entropy (BCE) and mean squared error (MSE). In this article, I will use the MNIST dataset for the demo. MNIST images have only one channel, and pixels take values between 0 and 1.
In this case, BCE loss can be used as reconstruction loss to treat pixels of MNIST images as a binary random variable that follows the Bernoulli distribution.
reconstruction_loss = nn.BCELoss(reduction='sum')
1.3 Kullback–Leibler Divergence
As mentioned above - KL divergence evaluates the difference between two distributions. Note that it does not have a symmetric property of a distance: KL(P‖Q)!=KL(Q‖P).
The two distributions that need to be compared are:
- the latent space of encoder output given input images x: q(z|x)
-
latent space prior p(z) which is assumed to be a normal distribution with a mean of zero and a standard deviation of one in each latent space dimension N(0, I).
Such an assumption simplifies the KL divergence computation and encourages the latent space to follow a known, manageable distribution.
from torch.distributions.kl import kl_divergence
def kl_divergence_loss(z_dist):
return kl_divergence(z_dist,
Normal(torch.zeros_like(z_dist.mean),
torch.ones_like(z_dist.stddev))
).sum(-1).sum()
1.4 Encoder
class Encoder(nn.Module):
def __init__(self, im_chan=1, output_chan=32, hidden_dim=16):
super(Encoder, self).__init__()
self.z_dim = output_chan
self.encoder = nn.Sequential(
self.init_conv_block(im_chan, hidden_dim),
self.init_conv_block(hidden_dim, hidden_dim * 2),
# double output_chan for mean and std with [output_chan] size
self.init_conv_block(hidden_dim * 2, output_chan * 2, final_layer=True),
)
def init_conv_block(self, input_channels, output_channels, kernel_size=4, stride=2, padding=0, final_layer=False):
layers = [
nn.Conv2d(input_channels, output_channels,
kernel_size=kernel_size,
padding=padding,
stride=stride)
]
if not final_layer:
layers += [
nn.BatchNorm2d(output_channels),
nn.ReLU(inplace=True)
]
return nn.Sequential(*layers)
def forward(self, image):
encoder_pred = self.encoder(image)
encoding = encoder_pred.view(len(encoder_pred), -1)
mean = encoding[:, :self.z_dim]
logvar = encoding[:, self.z_dim:]
# encoding output representing standard deviation is interpreted as
# the logarithm of the variance associated with the normal distribution
# take the exponent to convert it to standard deviation
return mean, torch.exp(logvar*0.5)
1.5 Decoder
class Decoder(nn.Module):
def __init__(self, z_dim=32, im_chan=1, hidden_dim=64):
super(Decoder, self).__init__()
self.z_dim = z_dim
self.decoder = nn.Sequential(
self.init_conv_block(z_dim, hidden_dim * 4),
self.init_conv_block(hidden_dim * 4, hidden_dim * 2, kernel_size=4, stride=1),
self.init_conv_block(hidden_dim * 2, hidden_dim),
self.init_conv_block(hidden_dim, im_chan, kernel_size=4, final_layer=True),
)
def init_conv_block(self, input_channels, output_channels, kernel_size=3, stride=2, padding=0, final_layer=False):
layers = [
nn.ConvTranspose2d(input_channels, output_channels,
kernel_size=kernel_size,
stride=stride, padding=padding)
]
if not final_layer:
layers += [
nn.BatchNorm2d(output_channels),
nn.ReLU(inplace=True)
]
else:
layers += [nn.Sigmoid()]
return nn.Sequential(*layers)
def forward(self, z):
# Ensure the input latent vector z is correctly reshaped for the decoder
x = z.view(-1, self.z_dim, 1, 1)
# Pass the reshaped input through the decoder network
return self.decoder(x)
1.6 VAE Model
To back-propagate through a random sample you need to move the parameters of the random sample (μ and 𝝈) outside of the the function to allow the gradient computation through the parameters. This step is also called the “reparameterization trick.”
In PyTorch, you can create a Normal distribution with the encoder’s output μ and 𝝈 and sample from it with rsample() method that implements the reparameterization trick: it is the same as torch.randn(z_dim) * stddev + mean)
class VAE(nn.Module):
def __init__(self, z_dim=32, im_chan=1):
super(VAE, self).__init__()
self.z_dim = z_dim
self.encoder = Encoder(im_chan, z_dim)
self.decoder = Decoder(z_dim, im_chan)
def forward(self, images):
z_dist = Normal(self.encoder(images))
# sample from distribution with reparametarazation trick
z = z_dist.rsample()
decoding = self.decoder(z)
return decoding, z_dist
1.7 Training a VAE
Load MNIST train and test data.
transform = transforms.Compose([transforms.ToTensor()])
# Download and load the MNIST training data
trainset = datasets.MNIST('.', download=True, train=True, transform=transform)
train_loader = DataLoader(trainset, batch_size=64, shuffle=True)
# Download and load the MNIST test data
testset = datasets.MNIST('.', download=True, train=False, transform=transform)
test_loader = DataLoader(testset, batch_size=64, shuffle=True)
VAE training steps
Create a training loop that follows the VAE training steps visualized in the figure above.
def train_model(epochs=10, z_dim = 16):
model = VAE(z_dim=z_dim).to(device)
model_opt = torch.optim.Adam(model.parameters())
for epoch in range(epochs):
print(f"Epoch {epoch}")
for images, step in tqdm(train_loader):
images = images.to(device)
model_opt.zero_grad()
recon_images, encoding = model(images)
loss = reconstruction_loss(recon_images, images)+ kl_divergence_loss(encoding)
loss.backward()
model_opt.step()
show_images_grid(images.cpu(), title=f'Input images')
show_images_grid(recon_images.cpu(), title=f'Reconstructed images')
return model
z_dim = 8
vae = train_model(epochs=20, z_dim=z_dim)
1.8 Visualize Latent Space
def visualize_latent_space(model, data_loader, device, method='TSNE', num_samples=10000):
model.eval()
latents = []
labels = []
with torch.no_grad():
for i, (data, label) in enumerate(data_loader):
if len(latents) > num_samples:
break
mu, _ = model.encoder(data.to(device))
latents.append(mu.cpu())
labels.append(label.cpu())
latents = torch.cat(latents, dim=0).numpy()
labels = torch.cat(labels, dim=0).numpy()
assert method in ['TSNE', 'UMAP'], 'method should be TSNE or UMAP'
if method == 'TSNE':
tsne = TSNE(n_components=2, verbose=1)
tsne_results = tsne.fit_transform(latents)
fig = px.scatter(tsne_results, x=0, y=1, color=labels, labels={'color': 'label'})
fig.update_layout(title='VAE Latent Space with TSNE',
width=600,
height=600)
elif method == 'UMAP':
reducer = umap.UMAP()
embedding = reducer.fit_transform(latents)
fig = px.scatter(embedding, x=0, y=1, color=labels, labels={'color': 'label'})
fig.update_layout(title='VAE Latent Space with UMAP',
width=600,
height=600
)
fig.show()
visualize_latent_space(vae, train_loader,
device='cuda' if torch.cuda.is_available() else 'cpu',
method='UMAP', num_samples=10000)
VAE model latent space visualized with UMAP
2 Sampling With VAE
Sampling from a Variational Autoencoder (VAE) enables the generation of new data that is similar to the one seen during training and it is a unique aspect that separates VAE from traditional AE architecture.
There are several ways of sampling from a VAE:
- posterior sampling: sampling from the posterior distribution given a provided input.
- prior sampling: sampling from the latent space assuming a standard normal multivariate distribution. This is possible due to the assumption (used during VAE training) that the latent variables are normally distributed. This method does not allow the generation of data with specific properties (for example, generating data from a specific class).
- interpolation: interpolation between two points in the latent space can reveal how changes in the latent space variable correspond to changes in the generated data.
-
traversal of latent dimensions: traversing latent dimensions of VAE latent space variance of the data depends on each dimension. Traversal is done by fixing all dimensions of the latent vector except one chosen dimension and varying values of the chosen dimension in its range. Some dimensions of the latent space may correspond to specific attributes of the data (VAE does not have specific mechanisms to force that behavior but it may happen).
For example, one dimension in latent space may control the emotional expression of a face or the orientation of an object.
Each sampling method provides a different way of exploring and understanding the data properties captured by the latent space of VAE.
2.1 Posterior Sampling (From a Given Input Image)
The encoder outputs a distribution (μ_x and 𝝈_x of normal distribution) in latent space. Sampling from normal distribution N(μ_x, 𝝈_x) and passing the sampled vector to the decoder results in generating an image similar to the given input image.
def posterior_sampling(model, data_loader, n_samples=25):
model.eval()
images, _ = next(iter(data_loader))
images = images[:n_samples]
with torch.no_grad():
_, encoding_dist = model(images.to(device))
input_sample=encoding_dist.sample()
recon_images = model.decoder(input_sample)
show_images_grid(images, title=f'input samples')
show_images_grid(recon_images, title=f'generated posterior samples')
posterior_sampling(vae, train_loader, n_samples=30)
Posterior sampling allows the generating of realistic data samples but with low variability: output data is similar to the input data.
2.2 Prior Sampling (From a Random Latent Space Vector)
Sampling from the distribution and passing the sampled vector to the decoder allows the generating of new data
def prior_sampling(model, z_dim=32, n_samples = 25):
model.eval()
input_sample=torch.randn(n_samples, z_dim).to(device)
with torch.no_grad():
sampled_images = model.decoder(input_sample)
show_images_grid(sampled_images, title=f'generated prior samples')
prior_sampling(vae, z_dim, n_samples=40)
Prior sampling with N(0, I) does not always generate plausible data but has high variability.
2.3 Sampling From Class Centers
Mean encodings of each class can be accumulated from the whole dataset and later be used for a controlled (conditional generation).
def get_data_predictions(model, data_loader):
model.eval()
latents_mean = []
latents_std = []
labels = []
with torch.no_grad():
for i, (data, label) in enumerate(data_loader):
mu, std = model.encoder(data.to(device))
latents_mean.append(mu.cpu())
latents_std.append(std.cpu())
labels.append(label.cpu())
latents_mean = torch.cat(latents_mean, dim=0)
latents_std = torch.cat(latents_std, dim=0)
labels = torch.cat(labels, dim=0)
return latents_mean, latents_std, labels
def get_classes_mean(class_to_idx, labels, latents_mean, latents_std):
classes_mean = {}
for class_name in train_loader.dataset.class_to_idx:
class_id = train_loader.dataset.class_to_idx[class_name]
labels_class = labels[labels==class_id]
latents_mean_class = latents_mean[labels==class_id]
latents_mean_class = latents_mean_class.mean(dim=0, keepdims=True)
latents_std_class = latents_std[labels==class_id]
latents_std_class = latents_std_class.mean(dim=0, keepdims=True)
classes_mean[class_id] = [latents_mean_class, latents_std_class]
return classes_mean
latents_mean, latents_stdvar, labels = get_data_predictions(vae, train_loader)
classes_mean = get_classes_mean(train_loader.dataset.class_to_idx, labels, latents_mean, latents_stdvar)
n_samples = 20
for class_id in classes_mean.keys():
latents_mean_class, latents_stddev_class = classes_mean[class_id]
# create normal distribution of the current class
class_dist = Normal(latents_mean_class, latents_stddev_class)
percentiles = torch.linspace(0.05, 0.95, n_samples)
# get samples from different parts of the distribution using icdf
# https://pytorch.org/docs/stable/distributions.html#torch.distributions.distribution.Distribution.icdf
class_z_sample = class_dist.icdf(percentiles[:, None].repeat(1, z_dim))
with torch.no_grad():
# generate image directly from mean
class_image_prototype = vae.decoder(latents_mean_class.to(device))
# generate images sampled from Normal(class mean, class std)
class_images = vae.decoder(class_z_sample.to(device))
show_image(class_image_prototype[0].cpu(), title=f'Class {class_id} prototype image')
show_images_grid(class_images.cpu(), title=f'Class {class_id} images')
Sampling from a normal distribution with averaged class μ guarantees the generation of new data from the same class.
image generated from class 3 center
image generated from class 4 center
Low and high percentile values used in icdf result in a high data variance
2.4 Interpolation
def linear_interpolation(start, end, steps):
# Create a linear path from start to end
z = torch.linspace(0, 1, steps)[:, None].to(device) * (end - start) + start
# Decode the samples along the path
vae.eval()
with torch.no_grad():
samples = vae.decoder(z)
return samples
2.4.1 Interpolation Between Two Random Latent Vectors
start = torch.randn(1, z_dim).to(device)
end = torch.randn(1, z_dim).to(device)
interpolated_samples = linear_interpolation(start, end, steps = 24)
show_images_grid(interpolated_samples, title=f'Linear interpolation between two random latent vectors')
2.4.2 Interpolation Between Two Class Centers
for start_class_id in range(1,10):
start = classes_mean[start_class_id][0].to(device)
for end_class_id in range(1, 10):
if end_class_id == start_class_id:
continue
end = classes_mean[end_class_id][0].to(device)
interpolated_samples = linear_interpolation(start, end, steps = 20)
show_images_grid(interpolated_samples, title=f'Linear interpolation between classes {start_class_id} and {end_class_id}')
2.5 Latent Space Traversal
Each dimension of the latent vector represents a normal distribution; the range of values of the dimension is controlled by mean and variance of the dimension. A simple way to traverse the range of values would be using inverse CDF (cumulative distribution functions) of the normal distribution.
ICDF takes a value between 0 and 1 (representing probability) and returns a value from the distribution. For a given probability p ICDF outputs a p_icdf value such that the probability of a random variable being <=p_icdf equals given probability p?”
If you have a normal distribution, icdf(0.5) should return the mean of the distribution. icdf(0.95) should return a value larger than 95% of the data from the distribution.
Visualization of CDF and values returned by ICDF given probabilities 0.025, 0.5, 0.975
2.5.1 Single Dimension Latent Space Traversal
def latent_space_traversal(model, input_sample, norm_dist, dim_to_traverse, n_samples, latent_dim, device):
# Create a range of values to traverse
assert input_sample.shape[0] == 1, 'input sample shape should be [1, latent_dim]'
# Generate linearly spaced percentiles between 0.05 and 0.95
percentiles = torch.linspace(0.1, 0.9, n_samples)
# Get the quantile values corresponding to the percentiles
traversed_values = norm_dist.icdf(percentiles[:, None].repeat(1, z_dim))
# Initialize a latent space vector with zeros
z = input_sample.repeat(n_samples, 1)
# Assign the traversed values to the specified dimension
z[:, dim_to_traverse] = traversed_values[:, dim_to_traverse]
# Decode the latent vectors
with torch.no_grad():
samples = model.decoder(z.to(device))
return samples
for class_id in range(0,10):
mu, std = classes_mean[class_id]
with torch.no_grad():
recon_images = vae.decoder(mu.to(device))
show_image(recon_images[0], title=f'class {class_id} mean sample')
for i in range(z_dim):
interpolated_samples = latent_space_traversal(vae, mu, norm_dist=Normal(mu, torch.ones_like(mu)), dim_to_traverse=i, n_samples=20, latent_dim=z_dim, device=device)
show_images_grid(interpolated_samples, title=f'Class {class_id} dim={i} traversal')
Traversing a single dimension may result in a change of digit style or control digit orientation.
2.5.3 Two Dimensions Latent Space Traversal
def traverse_two_latent_dimensions(model, input_sample, z_dist, n_samples=25, z_dim=16, dim_1=0, dim_2=1, title='plot'):
digit_size=28
percentiles = torch.linspace(0.10, 0.9, n_samples)
grid_x = z_dist.icdf(percentiles[:, None].repeat(1, z_dim))
grid_y = z_dist.icdf(percentiles[:, None].repeat(1, z_dim))
figure = np.zeros((digit_size * n_samples, digit_size * n_samples))
z_sample_def = input_sample.clone().detach()
# select two dimensions to vary (dim_1 and dim_2) and keep the rest fixed
for yi in range(n_samples):
for xi in range(n_samples):
with torch.no_grad():
z_sample = z_sample_def.clone().detach()
z_sample[:, dim_1] = grid_x[xi, dim_1]
z_sample[:, dim_2] = grid_y[yi, dim_2]
x_decoded = model.decoder(z_sample.to(device)).cpu()
digit = x_decoded[0].reshape(digit_size, digit_size)
figure[yi * digit_size: (yi + 1) * digit_size,
xi * digit_size: (xi + 1) * digit_size] = digit.numpy()
plt.figure(figsize=(6, 6))
plt.imshow(figure, cmap='Greys_r')
plt.title(title)
plt.show()
for class_id in range(10):
mu, std = classes_mean[class_id]
with torch.no_grad():
recon_images = vae.decoder(mu.to(device))
show_image(recon_images[0], title=f'class {class_id} mean sample')
traverse_two_latent_dimensions(vae, mu, z_dist=Normal(mu, torch.ones_like(mu)), n_samples=8, z_dim=z_dim, dim_1=3, dim_2=6, title=f'Class {class_id} traversing dimensions {(3, 6)}')
Traversing multiple dimensions at once provides a controllable way to generate data with high variability.
2.6 Bonus - 2D Manifold Of Digits From Latent Space
If a VAE model is trained with z_dim=2, it is possible to display a 2D manifold of digits from its latent space. To do that, I will use the traverse_two_latent_dimensions function with dim_1=0 and dim_2=2.
vae_2d = train_model(epochs=10, z_dim=2)
z_dist = Normal(torch.zeros(1, 2), torch.ones(1, 2))
input_sample = torch.zeros(1, 2)
with torch.no_grad():
decoding = vae_2d.decoder(input_sample.to(device))
traverse_two_latent_dimensions(vae_2d, input_sample, z_dist, n_samples=20, dim_1=0, dim_2=1, z_dim=2, title=f'traversing 2D latent space')
2D latent space